Documentation Index
Fetch the complete documentation index at: https://bloodhound.specterops.io/llms.txt
Use this file to discover all available pages before exploring further.
Before you begin
Full OpenGraph support requires a PostgreSQL graph database and one of the following editions:- BloodHound Enterprise (uses PostgreSQL by default)
-
BloodHound Community v8.0.0+ (requires changing to a PostgreSQL database)
While many OpenGraph features may work on a Neo4j database, there are functional and performance limitations (see the OpenGraph FAQ). For full support, migrate to a PostgreSQL database.
File requirements
OpenGraph data payloads are accepted as.json files or .zip archives. A single upload operation can include a mix of valid .json and .zip files from different collectors (for example, SharpHound and OpenGraph). Additional constraints include:
.ziparchives can contain multiple.jsonpayload files.- Nested
.ziparchives are not supported. - If you upload a
.ziparchive, every file in the archive must be a valid data payload.
Data payload structure
All OpenGraph extensions must use the same data payload format. At minimum, a data payload must include the following top-level structure:nodes and edges arrays must conform to minimum JSON schemas. BloodHound validates that the JSON is well-formed and that nodes and edges meet these schema requirements, but it does not enforce additional structure or constraints beyond them.
For field-level details and schema requirements, see the following pages:
| Page | Description |
|---|---|
| Metadata | Defines payload-wide ingest behavior to facilitate organizing and managing graph data by source. |
| Nodes | Represents entities in your graph, such as users, devices, repositories, applications, and environments. |
| Edges | Represents relationships between nodes in your graph indicating some form of interaction or access. |
Graph structure
To enforce additional structure beyond the minimum JSON schemas and enable advanced features in BloodHound, you can use an to define node and edge types, environments, and relationships. When you upload a data payload that conforms to an installed extension definition schema, BloodHound produces a structured graph with enhanced features compared to a generic graph.| Graph | Use this when |
|---|---|
| You only need to ingest and search nodes and edges. | |
| You need schema-driven analysis and pathfinding. |
Data source
OpenGraph extensions can collect data from many different systems. By default, BloodHound identifies this as sourceless data. If you’re using multiple extensions, you may have multiple data sources feeding into your BloodHound graph database without any way to differentiate them. To facilitate organizing and managing OpenGraph data in BloodHound, you can register asource_kind and apply it to the relevant nodes in your data payload. Once registered, BloodHound reuses it for subsequent uploads with the same node kinds, even if the data payload omits it.
For example, if you wanted to delete all data for a source, you could navigate to the Administration > Database Management page and select the name of the defined source.
You can still remove OpenGraph data without registering and applying a
source_kind by checking the Sourceless data option. However, that could have unintended consequences if you have multiple OpenGraph data sources without a registered source_kind.source_kind in two ways:
- Specify it in at least one payload’s
metadataobject. After that initial registration, later payloads can reuse it without re-registering it. - Define it as an
environments.source_kindvalue in an extension definition schema.
source_kind in two ways:
- Specify it in the payload’s
metadataobject, which applies it to all defined nodes. - Explicitly include it in a node’s
kindsarray, which applies it only to that node.
source_kind to produce and metrics.
The following points describe how source_kind influences metrics calculations:
- Exposure metrics: Traverse upstream toward the origin and stop when a node in the same zone is encountered (for Tier Zero paths, this means stopping when a Tier Zero node is reached). The origin node itself is not included.
- Impact metrics: Traverse downstream from the target node and include nodes in the same zone until the downstream set is fully enumerated.
Minimal viable data payload
The following is a minimal example payload for a generic graph. You can use it as a starting point for your own OpenGraph data payload. Copy and paste the following example into a new.json file or download this example file.
Run:
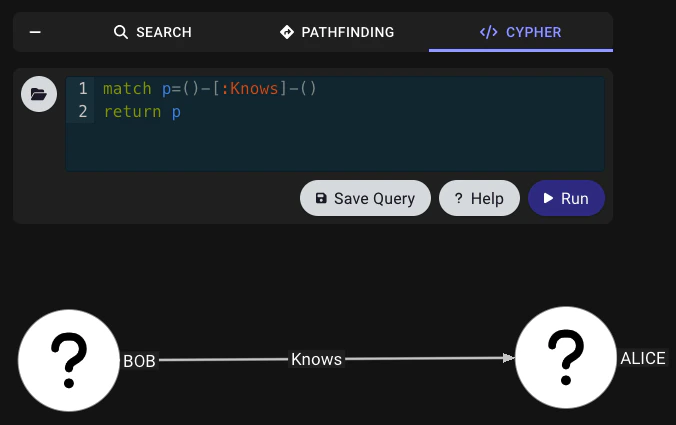
Troubleshooting
- Upload fails before schema checks: Confirm the file is valid JSON and encoded in UTF-8.
-
Upload fails with payload structure errors: Confirm the payload has top-level
graphwith bothnodesandedgesarrays. - ZIP upload fails unexpectedly: Ensure the archive does not contain nested ZIP files.